| RIKEN Center for Developmental Biology (CDB) 2-2-3 Minatojima minamimachi, Chuo-ku, Kobe 650-0047, Japan |
| XsalF comes to the fore in brain regionalization | ||||
July 13, 2004 - In vertebrates, the nervous system is divided into distinct regions patterned in a head-to-tail direction. Embryologists have long been interested in working out the means by which this regionalization is achieved, and for years the dominant theory has involved a two-step mechanism in which neural tissues are first induced lengthwise down the entire body axis, followed by a transformation step where a second signal specifies the posterior identities of the neural tissue, which can be induced by a number of factors including members of the Wnt and FGF gene families, and retinoic acid. While there is much evidence to support this model, a number of recent studies have indicated
that the specification of the forebrain (an anterior structure, presumed to be induced by the
initial signal in the activation-transformation model) requires additional regulatory inputs as
well. New work by scientists in the Laboratory for
Organogenesis and Neurogenesis (Group Director,
Yoshiki Sasai) at the RIKEN Center for Developmental Biology (CDB; Kobe, Japan) showing anterior
neural specifying activity in the African clawed frog, Xenopus laevis , lends weight
to the revisionist argument.
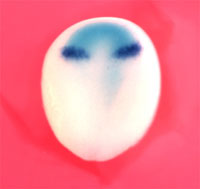
In an article published in the July 13 issue of Developmental Cell , Takayuki Onai et al. report that XsalF , the Xenopus homolog of spalt , a homeotic gene known to function in anterior-posterior segment identity in Drosophila , regulates the expression of forebrain and midbrain-specific genes. A series of experiments in which XsalF was misexpressed, deleted and its function blocked, showed direct linkage between XsalF expression and forebrain/midbrain identity. XsalF was originally identified in a screen of the frog anterior neural plate, a structure
that appears early in neural development. Sequencing of the gene, and analysis of the timing and
spatial pattern of its expression showed that it codes a transcription factor related to Spalt
and is expressed in the incipient forebrain and midbrain at precisely those developmental stages
when brain regions are specified. When members of the Sasai lab overexpressed the gene by injecting
its mRNA into early embryos, they found it caused the expanded expression of genes specific to
anterior brain regions while suppressing the expression of more posterior markers.
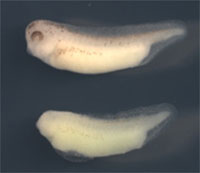
Onai et al. next went on to test the effects of the loss of XsalF function. Disabling the gene by the deletion of functional domains resulted in embryos with significant reductions in anterior neural structures. Conditional loss-of-function experiments showed that the gene's expression was required in the early and mid neurula stages, when the developing brain undergoes regionalization. They then confirmed the specificity of this requirement for XsalF by injecting morpholinos (short nucleotide chains that block the function of a targeted gene), which gave similar results to the earlier loss-of-function studies. These preliminary findings prompted the lab to look into the molecular mechanisms behind XsalF 's anterior specifying activity, which they suspected was linked to the inhibition of the Wnt cascade (a signaling pathway that posteriorizes neural tissues). They focused on two factors, GSK3ß and Tcf3 , known to antagonize Wnt signaling in certain contexts, and found that the expression of both factors was dependent on XsalF . Gain- and loss-of-function studies reconfirmed the connection between XsalF and these Wnt antagonists, showing that XsalF alters the receptivity of anterior neural cells to Wnt signaling by regulating the expression of GSK3ß and Tcf3 , making these cells resistant to the posteriorizing effects of Wnt. The signaling networks at work in this competitive regional determination appear to be intricate
and involved in two linked, but distinct, aspects of regions-specific transcriptional regulation - the
switching on of fore- and midbrain-specific genes, and the suppression of posterior genes. The
Sasai lab now plans to conduct further studies to develop a detailed model for understanding both
the direct and mediated effects of XsalF in neural patterning. |
||||
|
||||
[ Contact ] Douglas Sipp : sipp@cdb.riken.jp TEL : +81-78-306-3043 RIKEN CDB, Office for Science Communications and International Affairs |
| Copyright (C) CENTER FOR DEVELOPMENTAL BIOLOGY All rights reserved. |