| RIKEN Center for Developmental Biology (CDB) 2-2-3 Minatojima minamimachi, Chuo-ku, Kobe 650-0047, Japan |
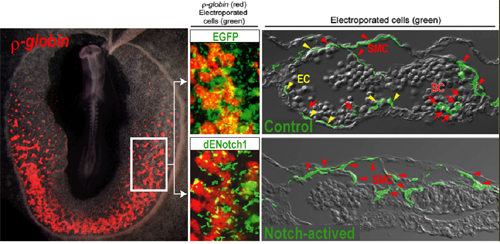
January 30, 2009 Not all development takes places inside the embryo. Many important functions, such as the formation of blood and blood vessels, are relegated to extraembryonic tissues in early developmental stages and reintroduced to the embryo proper subsequently. In the chicken embryo, the extraembryonic mesoderm gives rise to both blood and endothelial cells, but it has been never been shown whether this represents its full differentiative gamut. Now, a study by Masahiro Shin and Hiroki Nagai in the Laboratory for Early Embryogenesis (Guojun Sheng; Team Leader) reveals that this same extraembryonic tissue is the earliest source of smooth muscle cells as well. In an article published in Development, the team identified the transcription factor dHand as a genetic marker of smooth muscle and showed that the segregation of this population from blood and endothelial progenitors is mediated by Notch signaling. Notch has been shown to be involved in vascular remodeling and specification, but as little was known about its role in ventral mesodermal differentiation (which gives rise to the population of cells that populate the extraembryonic mesoderm), Shin made a truncated Notch construct engineered such that the pathway was continuously switched on and introduced it into primitive streak-stage embryos to study the effects on cell fate. He found that a preponderance of the Notch-expressing cells took on a smooth muscle fate, with very few differentiating into blood/endothelial progenitors. In situ hybridization revealed that Notch is active in the posterior primitive streak, where ventral mesoderm is generated.
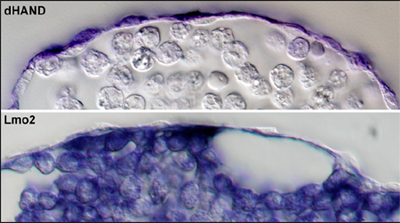
Conjecturing that Notch might act as a sorting mechanism to separate smooth muscle from blood and vascular progenitors, the team next sought a means of labeling smooth muscle, as no genetic markers for these cells had been identified. An in situ screen turned up the transcriptional factor dHand, which they found to be expressed in early smooth muscle cells as well as vascular smooth muscle layer and extraembryonic somatic mesoderm (also known to consist of smooth muscle cells) at later stages of development. Blood and endothelial progenitors have their own specific marker in the gene Scl. When the team went back and looked at expression patterns in the constitutively Notch active posterior streak, they found that cells expressing Notch also tended to express dHand, and not Scl. Inhibiting the Notch pathway abolished this exclusion. Interestingly, they found that although Notch activation is associated with nearly exclusive smooth muscle cell contribution to the extraembryonic mesoderm, the absence of Notch has no effect on the induction or differentiation of these cells. They asked next whether other signaling pathways known to work in the same vicinity might also be involved in smooth muscle specification. While BMP appeared to function primarily by ventralizing the mesoderm (leading to the induction of both smooth muscle and blood/endothelium), ectopic expression of Wnt strongly induced the smooth muscle marker dHand. Intriguingly, the signal responsible for inducing the blood/endothelial common progenitor has yet to be identified.
"Perhaps many researchers aren't so interested in extraembryonic tissues, but a similar tug-of-war between muscles and vessels happens in cardiac development and in mesoderm stem cell differentiation in vitro," says Sheng. "So in that sense the molecular mechanisms uncovered in our studies may be of more general interest." |
|||||||||
|
|||||||||
 |
| Copyright (C) CENTER FOR DEVELOPMENTAL BIOLOGY All rights reserved. |