| RIKEN Center for Developmental Biology (CDB) 2-2-3 Minatojima minamimachi, Chuo-ku, Kobe 650-0047, Japan |
December 4, 2010 Powerful molecular players frequently require major powerful inhibitors to keep them in check. Bone morphogenetic protein, or BMP, is one such factor, playing central roles in multiple developmental processes; its suppression in appriopriate contexts is likewise a critical component of many embryogenetic programs. The induction and patterning of neural tissues provides a good example of this to and fro. BMP, which is highly expressed on the embryos ventral side, is suppressed by inhibitors secreted from the Spemann organizer, leading to neural induction. BMP signaling is also involved in the subsequent pattern formation in this neural tissue. But, while it is clear that the suppression of the BMP pathway at given sites and timepoints is crucial to the patterning of the early embryo, just how this is accomplised is imperfectly understood.
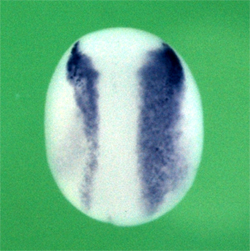
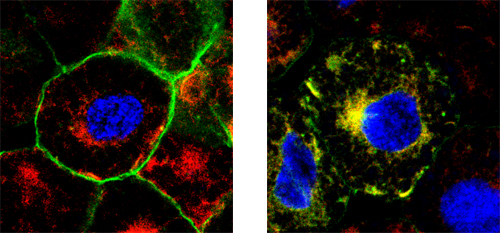
Toshihiro Aramaki and colleagues in the Laboratory for Organogenesis and Neurogenesis (Yoshiki Sasai, Group Director) have shed some light on this process through the lens of a new controller of BMP signaling. In a report published in Developmental Cell, the group identifies the membrane protein Jiraiya as a specific inhibitor of a subunit of the BMP receptor in the African clawed frog, Xenopus laevis. Jiraiya first caught the researchers eye when it was captured in a screen for genes downstream of the BMP inhibitor Chordin. Its dorsal expression pattern recalled to mind a character, named Jiraiya, who appears mounted on an amphibian in a famous kaubuki play, thus earning it its colorful name. The protein is first expressed in the dorsal ectoderm of the early gastrula, shifting subsequently to the dorsal aspect of the neural tube. When Aramaki knocked down its function using a morpholino antisense oligonucleotide, he saw an increase in the scope of expression of lateral neural plate marker genes in the treated embryos, accompanied by upregulation of activated Smad1, a factor downstream in the BMP pathway, suggesting that this newly identified protein worked to limit dorsal neuralization. To get a clearer sense of the mechanism, the group used an animal cap assay of embryonic cells to test the effect of Jiraiya overexpression. They found that Jiraiya interferes with the function of BMPRII, one of two subunits of the BMP receptor. The suppressor effect was specific to BMP; no inihibition was seen, for example, of a second member of the TGF-ß family, activin. The question was: how does Jiraiya achieve this? Immunostaining revealed that Jiraiya tended to accumulate more in the cytoplasm than at the plasma membrane, suggesting it might work intracellularly to block the function of the membrane-bound BMPRII, which was supported by the failure of this receptor component to integrate into the cell membrane in cells overexpressing Jiraiya. They next looked for the site of interaction between these proteins by creating a series of mutant proteins from which various domains were deleted. They found that when BMPRII lost its C-terminal, it became much less sensitive to Jiraiyas quelling effect. Compellingly, the BMPRII tail domain contains a highly conserved 6-amino snippet not found in any other TGF-ß receptor family member, suggesting a locus for the specifity of the BMPRII-Jiraiya interaction. The finding represents the first identification of a receptor-level inhibitor of BMP.
We hope to be able to tease out Jiraiyas mechanism of action in vivo, says Sasai. It will be especially interesting to work out how Jiraiya functions in mammals, and specifically whether it inhibits the BMP pathway there as well. |
|||||||
|
|||||||
 |
| Copyright (C) CENTER FOR DEVELOPMENTAL BIOLOGY All rights reserved. |