| RIKEN Center for Developmental Biology (CDB) 2-2-3 Minatojima minamimachi, Chuo-ku, Kobe 650-0047, Japan |
January 17, 2011 As in manmade timepieces, the movements of the genetic clockworks that lie behind circadian cycles involve a remarkable amount of complexity. The mammalian circadian clock, for example, is thought to arise from the interactions of around 20 transcription factors with specific DNA sequences associated with morning, day, and night expression. Existing models of this genetic network can readily explain the basis for the day and night activities, but the mechanism underlying morning expression remains incompletely understood. It is thought that delayed negative feedback exerted by the morning (E/E? box) inhibitor Cryptochrome 1 (Cry1), which is itself expressed in evening, plays an important role in keeping the biological clock on time. But just how it achieves this effect is unknown. Maki Ukai-Tadenuma and Rikuhiro G. Yamada of the Laboratory for Systems Biology (Hiroki R. Ueda, Project Leader), along with colleagues in the Universities of Memphis (USA) and Fribourg (Switzerland), now report how delayed feedback repression is a key factor in mammalian clock function. Published in Cell, this work shows the role of Cry1 as mediator of delayed negative feedback repression and fleshes out the current understanding of the circadian circuitry.
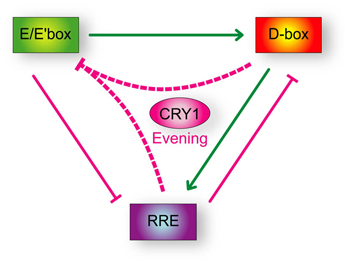
The team began by looking into the basis for the evening expression of Cry1 using reporter genes coding for the luciferase protein to detect transcriptional activity, and found that the Cry1 promoter region induces the expression of genes carrying the daytime expression motif. A closer look at Cry1s DNA revealed that its intronic region contains a separate sequence that induces nighttime clock genes. They next stitched together a construct including these promoter and intron regions, and ran another reporter assay to observe its behavior, and found that its expression switched on in circadian evening, suggesting that this in-between expression time is a result of the combination of day and night regulatory elements. To test this model, the team tried to rescue clock function in cells with homozygous deletions of both Cry1 and Cry2 by inducing the evening expression of exogenous Cry1. They found that while the Cry1 promoter region alone was ineffective, when the promoter and intron regions were used in conjunction, the genes circadian rhythmicity was restored. Using this same set-up, Ukai-Tadenuma and Yamada next tried changing the onset time of Cry1 expression, and found that as expression neared midday, meaning that the normal phase delay was reduced, the amplitude of circadian oscillations grew smaller, in line with predictions. Similarly, prolonging the delay of exogenous Cry1 expression caused an increase in the length of the restored cycle. The team's findings were recapitulated by a relatively simple phase vector model, which not only successfully reproduces the findings from the current study, but numerous other aspects of the circadian clock network as well. "In 1990, Paul Hardin at Texas A&M pointed out the importance of delayed feedback repression in biological clocks, but it has taken 21 years to work out the mechanism behind it," says Ueda. "We will continue exploring whether the current minimal transcriptional network model is complete, or whether new regulatory systems remain to be discovered." |
|||||
|
|||||
 |
| Copyright (C) CENTER FOR DEVELOPMENTAL BIOLOGY All rights reserved. |